| Rank | Motif | Name | P-value | log P-pvalue | q-value (Benjamini) | # Target Sequences with Motif | % of Targets Sequences with Motif | # Background Sequences with Motif | % of Background Sequences with Motif | Motif File |
PDF |
| 1 |  | Lhx3(Homeobox)/Neuron-Lhx3-ChIP-Seq(GSE31456)/Homer | 1e-12 | -2.934e+01 | 0.0000 | 80.0 | 51.28% | 11293.6 | 24.00% | motif file (matrix) |
pdf |
| 2 |  | PAX5(Paired/Homeobox)/GM12878-PAX5-ChIP-Seq(GSE32465)/Homer | 1e-11 | -2.737e+01 | 0.0000 | 22.0 | 14.10% | 951.7 | 2.02% | motif file (matrix) |
pdf |
| 3 |  | Lhx2(Homeobox)/HFSC-Lhx2-ChIP-Seq(GSE48068)/Homer | 1e-11 | -2.558e+01 | 0.0000 | 59.0 | 37.82% | 7235.8 | 15.38% | motif file (matrix) |
pdf |
| 4 |  | PAX5/long/HOCOMOCO | 1e-10 | -2.517e+01 | 0.0000 | 16.0 | 10.26% | 488.0 | 1.04% | motif file (matrix) |
pdf |
| 5 |  | Nkx6.1(Homeobox)/Islet-Nkx6.1-ChIP-Seq(GSE40975)/Homer | 1e-9 | -2.235e+01 | 0.0000 | 98.0 | 62.82% | 17747.1 | 37.72% | motif file (matrix) |
pdf |
| 6 |  | FOXA1(Forkhead)/MCF7-FOXA1-ChIP-Seq(GSE26831)/Homer | 1e-9 | -2.124e+01 | 0.0000 | 48.0 | 30.77% | 5708.1 | 12.13% | motif file (matrix) |
pdf |
| 7 |  | FOXA1(Forkhead)/LNCAP-FOXA1-ChIP-Seq(GSE27824)/Homer | 1e-8 | -1.994e+01 | 0.0000 | 53.0 | 33.97% | 6991.3 | 14.86% | motif file (matrix) |
pdf |
| 8 |  | Fox:Ebox(Forkhead:HLH)/Panc1-Foxa2-ChIP-Seq(GSE47459)/Homer | 1e-8 | -1.911e+01 | 0.0000 | 42.0 | 26.92% | 4872.9 | 10.36% | motif file (matrix) |
pdf |
| 9 |  | Isl1(Homeobox)/Neuron-Isl1-ChIP-Seq(GSE31456)/Homer | 1e-8 | -1.906e+01 | 0.0000 | 70.0 | 44.87% | 11140.1 | 23.68% | motif file (matrix) |
pdf |
| 10 |  | Nanog(Homeobox)/mES-Nanog-ChIP-Seq(GSE11724)/Homer | 1e-7 | -1.769e+01 | 0.0000 | 108.0 | 69.23% | 22187.9 | 47.16% | motif file (matrix) |
pdf |
| 11 |  | PAX6/SwissRegulon | 1e-7 | -1.764e+01 | 0.0000 | 17.0 | 10.90% | 945.7 | 2.01% | motif file (matrix) |
pdf |
| 12 |  | Pax8(Paired/Homeobox)/Thyroid-Pax8-ChIP-Seq(GSE26938)/Homer | 1e-7 | -1.762e+01 | 0.0000 | 15.0 | 9.62% | 717.9 | 1.53% | motif file (matrix) |
pdf |
| 13 |  | NeuroD1(bHLH)/Islet-NeuroD1-ChIP-Seq(GSE30298)/Homer | 1e-7 | -1.670e+01 | 0.0000 | 30.0 | 19.23% | 3000.7 | 6.38% | motif file (matrix) |
pdf |
| 14 |  | MA0069.1_PAX6/Jaspar | 1e-7 | -1.631e+01 | 0.0000 | 12.0 | 7.69% | 475.1 | 1.01% | motif file (matrix) |
pdf |
| 15 |  | HOXD13(Homeobox)/Chicken-Hoxd13-ChIP-Seq(GSE38910)/Homer | 1e-6 | -1.437e+01 | 0.0000 | 47.0 | 30.13% | 6866.7 | 14.60% | motif file (matrix) |
pdf |
| 16 |  | Foxa2(Forkhead)/Liver-Foxa2-ChIP-Seq(GSE25694)/Homer | 1e-6 | -1.428e+01 | 0.0000 | 35.0 | 22.44% | 4332.6 | 9.21% | motif file (matrix) |
pdf |
| 17 |  | PHA-4(Forkhead)/cElegans-Embryos-PHA4-ChIP-Seq(modEncode)/Homer | 1e-5 | -1.369e+01 | 0.0000 | 88.0 | 56.41% | 17600.5 | 37.41% | motif file (matrix) |
pdf |
| 18 |  | Pdx1(Homeobox)/Islet-Pdx1-ChIP-Seq(SRA008281)/Homer | 1e-5 | -1.168e+01 | 0.0002 | 40.0 | 25.64% | 5952.6 | 12.65% | motif file (matrix) |
pdf |
| 19 |  | LIN-39(Homeobox)/cElegans.L3-LIN39-ChIP-Seq(modEncode)/Homer | 1e-4 | -1.148e+01 | 0.0002 | 48.0 | 30.77% | 7850.8 | 16.69% | motif file (matrix) |
pdf |
| 20 |  | PAX6/HOCOMOCO | 1e-4 | -1.114e+01 | 0.0002 | 9.0 | 5.77% | 425.3 | 0.90% | motif file (matrix) |
pdf |
| 21 |  | Olig2(bHLH)/Neuron-Olig2-ChIP-Seq(GSE30882)/Homer | 1e-4 | -1.080e+01 | 0.0003 | 50.0 | 32.05% | 8534.0 | 18.14% | motif file (matrix) |
pdf |
| 22 |  | Foxo1(Forkhead)/RAW-Foxo1-ChIP-Seq(Fan et al.)/Homer | 1e-4 | -1.073e+01 | 0.0003 | 50.0 | 32.05% | 8557.7 | 18.19% | motif file (matrix) |
pdf |
| 23 |  | Nkx2.1(Homeobox)/LungAC-Nkx2.1-ChIP-Seq(GSE43252)/Homer | 1e-4 | -1.058e+01 | 0.0004 | 62.0 | 39.74% | 11635.1 | 24.73% | motif file (matrix) |
pdf |
| 24 |  | Rfx5(HTH)/GM12878-Rfx5-ChIP-Seq(GSE31477)/Homer | 1e-4 | -9.229e+00 | 0.0013 | 13.0 | 8.33% | 1114.8 | 2.37% | motif file (matrix) |
pdf |
| 25 |  | Ptf1a(HLH)/Panc1-Ptf1a-ChIP-Seq(GSE47459)/Homer | 1e-3 | -8.974e+00 | 0.0016 | 54.0 | 34.62% | 10158.3 | 21.59% | motif file (matrix) |
pdf |
| 26 |  | PAX3:FKHR-fusion(Paired/Homeobox)/Rh4-PAX3:FKHR-ChIP-Seq(GSE19063)/Homer | 1e-3 | -8.880e+00 | 0.0017 | 12.0 | 7.69% | 998.1 | 2.12% | motif file (matrix) |
pdf |
| 27 |  | EGL-5(Homeobox)/cElegans-L3-EGL5-ChIP-Seq(modEncode)/Homer | 1e-3 | -8.009e+00 | 0.0040 | 50.0 | 32.05% | 9503.3 | 20.20% | motif file (matrix) |
pdf |
| 28 |  | BMYB(HTH)/Hela-BMYB-ChIPSeq(GSE27030)/Homer | 1e-3 | -7.850e+00 | 0.0045 | 41.0 | 26.28% | 7312.3 | 15.54% | motif file (matrix) |
pdf |
| 29 |  | E2A(HLH)/proBcell-E2A-ChIP-Seq(GSE21978)/Homer | 1e-3 | -7.835e+00 | 0.0045 | 31.0 | 19.87% | 4955.7 | 10.53% | motif file (matrix) |
pdf |
| 30 |  | PDX1/HOCOMOCO | 1e-3 | -7.809e+00 | 0.0045 | 21.0 | 13.46% | 2799.0 | 5.95% | motif file (matrix) |
pdf |
| 31 |  | Nkx2.5(Homeobox)/HL1-Nkx2.5.biotin-ChIP-Seq(GSE21529)/Homer | 1e-3 | -7.783e+00 | 0.0045 | 49.0 | 31.41% | 9336.8 | 19.85% | motif file (matrix) |
pdf |
| 32 |  | AMYB(HTH)/Testes-AMYB-ChIP-Seq(GSE44588)/Homer | 1e-3 | -7.708e+00 | 0.0045 | 40.0 | 25.64% | 7118.7 | 15.13% | motif file (matrix) |
pdf |
| 33 |  | Eomes(T-box)/H9-Eomes-ChIP-Seq(GSE26097)/Homer | 1e-3 | -7.446e+00 | 0.0057 | 55.0 | 35.26% | 11033.3 | 23.45% | motif file (matrix) |
pdf |
| 34 |  | Ap4(HLH)/AML-Tfap4-ChIP-Seq(GSE45738)/Homer | 1e-3 | -7.134e+00 | 0.0076 | 29.0 | 18.59% | 4705.1 | 10.00% | motif file (matrix) |
pdf |
| 35 |  | RFX(HTH)/K562-RFX3-ChIP-Seq(SRA012198)/Homer | 1e-3 | -7.095e+00 | 0.0077 | 5.0 | 3.21% | 216.2 | 0.46% | motif file (matrix) |
pdf |
| 36 |  | MyoG(HLH)/C2C12-MyoG-ChIP-Seq(GSE36024)/Homer | 1e-3 | -6.957e+00 | 0.0085 | 25.0 | 16.03% | 3856.7 | 8.20% | motif file (matrix) |
pdf |
| 37 |  | Atoh1(bHLH)/Cerebellum-Atoh1-ChIP-Seq(GSE22111)/Homer | 1e-3 | -6.941e+00 | 0.0085 | 26.0 | 16.67% | 4083.8 | 8.68% | motif file (matrix) |
pdf |
| 38 |  | Rfx1(HTH)/NPC-H3K4me1-ChIP-Seq(GSE16256)/Homer | 1e-3 | -6.909e+00 | 0.0085 | 8.0 | 5.13% | 604.7 | 1.29% | motif file (matrix) |
pdf |
| 39 |  | ISL1/HOCOMOCO | 1e-2 | -6.881e+00 | 0.0085 | 39.0 | 25.00% | 7172.9 | 15.25% | motif file (matrix) |
pdf |
| 40 |  | bZIP:IRF/Th17-BatF-ChIP-Seq(GSE39756)/Homer | 1e-2 | -6.664e+00 | 0.0103 | 16.0 | 10.26% | 2036.0 | 4.33% | motif file (matrix) |
pdf |
| 41 |  | Myf5(bHLH)/GM-Myf5-ChIP-Seq(GSE24852)/Homer | 1e-2 | -6.547e+00 | 0.0113 | 19.0 | 12.18% | 2669.0 | 5.67% | motif file (matrix) |
pdf |
| 42 |  | HNF6(Homeobox)/Liver-Hnf6-ChIP-Seq(ERP000394)/Homer | 1e-2 | -6.498e+00 | 0.0116 | 20.0 | 12.82% | 2891.0 | 6.14% | motif file (matrix) |
pdf |
| 43 |  | HOXA9(Homeobox)/HSC-Hoxa9-ChIP-Seq(GSE33509)/Homer | 1e-2 | -6.438e+00 | 0.0120 | 25.0 | 16.03% | 4003.2 | 8.51% | motif file (matrix) |
pdf |
| 44 |  | Rfx2(HTH)/LoVo-RFX2-ChIP-Seq(GSE49402)/Homer | 1e-2 | -6.169e+00 | 0.0154 | 5.0 | 3.21% | 267.7 | 0.57% | motif file (matrix) |
pdf |
| 45 |  | HLH-1(bHLH)/cElegans-Embryo-HLH1-ChIP-Seq(modEncode)/Homer | 1e-2 | -6.068e+00 | 0.0166 | 22.0 | 14.10% | 3438.3 | 7.31% | motif file (matrix) |
pdf |
| 46 |  | Unknown(Homeobox)/Limb-p300-ChIP-Seq/Homer | 1e-2 | -5.658e+00 | 0.0245 | 24.0 | 15.38% | 4013.6 | 8.53% | motif file (matrix) |
pdf |
| 47 |  | Hoxb4(Homeobox)/ES-Hoxb4-ChIP-Seq(GSE34014)/Homer | 1e-2 | -5.617e+00 | 0.0250 | 10.0 | 6.41% | 1089.7 | 2.32% | motif file (matrix) |
pdf |
| 48 |  | Tbx5(T-box)/HL1-Tbx5.biotin-ChIP-Seq(GSE21529)/Homer | 1e-2 | -5.567e+00 | 0.0257 | 60.0 | 38.46% | 13303.9 | 28.28% | motif file (matrix) |
pdf |
| 49 |  | Tcf12(HLH)/GM12878-Tcf12-ChIP-Seq(GSE32465)/Homer | 1e-2 | -5.303e+00 | 0.0328 | 21.0 | 13.46% | 3438.3 | 7.31% | motif file (matrix) |
pdf |
| 50 |  | RFX1..5_RFXANK_RFXAP.p2/SwissRegulon | 1e-2 | -5.284e+00 | 0.0328 | 7.0 | 4.49% | 624.6 | 1.33% | motif file (matrix) |
pdf |
| 51 |  | Hoxc9(Homeobox)/Ainv15-Hoxc9-ChIP-Seq(GSE21812)/Homer | 1e-2 | -5.206e+00 | 0.0347 | 19.0 | 12.18% | 3017.9 | 6.41% | motif file (matrix) |
pdf |
| 52 | 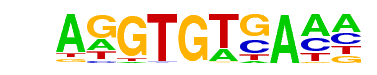 | Tbet(T-box)/CD8-Tbet-ChIP-Seq(GSE33802)/Homer | 1e-2 | -5.000e+00 | 0.0419 | 28.0 | 17.95% | 5195.6 | 11.04% | motif file (matrix) |
pdf |
| 53 |  | HOXA2(Homeobox)/mES-Hoxa2-ChIP-Seq(Donaldson et al.)/Homer | 1e-2 | -4.920e+00 | 0.0445 | 6.0 | 3.85% | 508.3 | 1.08% | motif file (matrix) |
pdf |
| 54 |  | Unknown3/Arabidopsis-Promoters/Homer | 1e-2 | -4.841e+00 | 0.0473 | 5.0 | 3.21% | 367.5 | 0.78% | motif file (matrix) |
pdf |
| 55 |  | X-box(HTH)/NPC-H3K4me1-ChIP-Seq(GSE16256)/Homer | 1e-2 | -4.786e+00 | 0.0490 | 5.0 | 3.21% | 372.4 | 0.79% | motif file (matrix) |
pdf |
| 56 |  | MyoD(HLH)/Myotube-MyoD-ChIP-Seq(GSE21614)/Homer | 1e-2 | -4.746e+00 | 0.0501 | 17.0 | 10.90% | 2705.3 | 5.75% | motif file (matrix) |
pdf |
| 57 |  | Pax6-mouse/UniProbe | 1e-2 | -4.697e+00 | 0.0517 | 19.0 | 12.18% | 3171.8 | 6.74% | motif file (matrix) |
pdf |
| 58 |  | Nkx3.1(Homeobox)/LNCaP-Nkx3.1-ChIP-Seq(GSE28264)/Homer | 1e-2 | -4.615e+00 | 0.0551 | 47.0 | 30.13% | 10283.3 | 21.86% | motif file (matrix) |
pdf |



















































































































